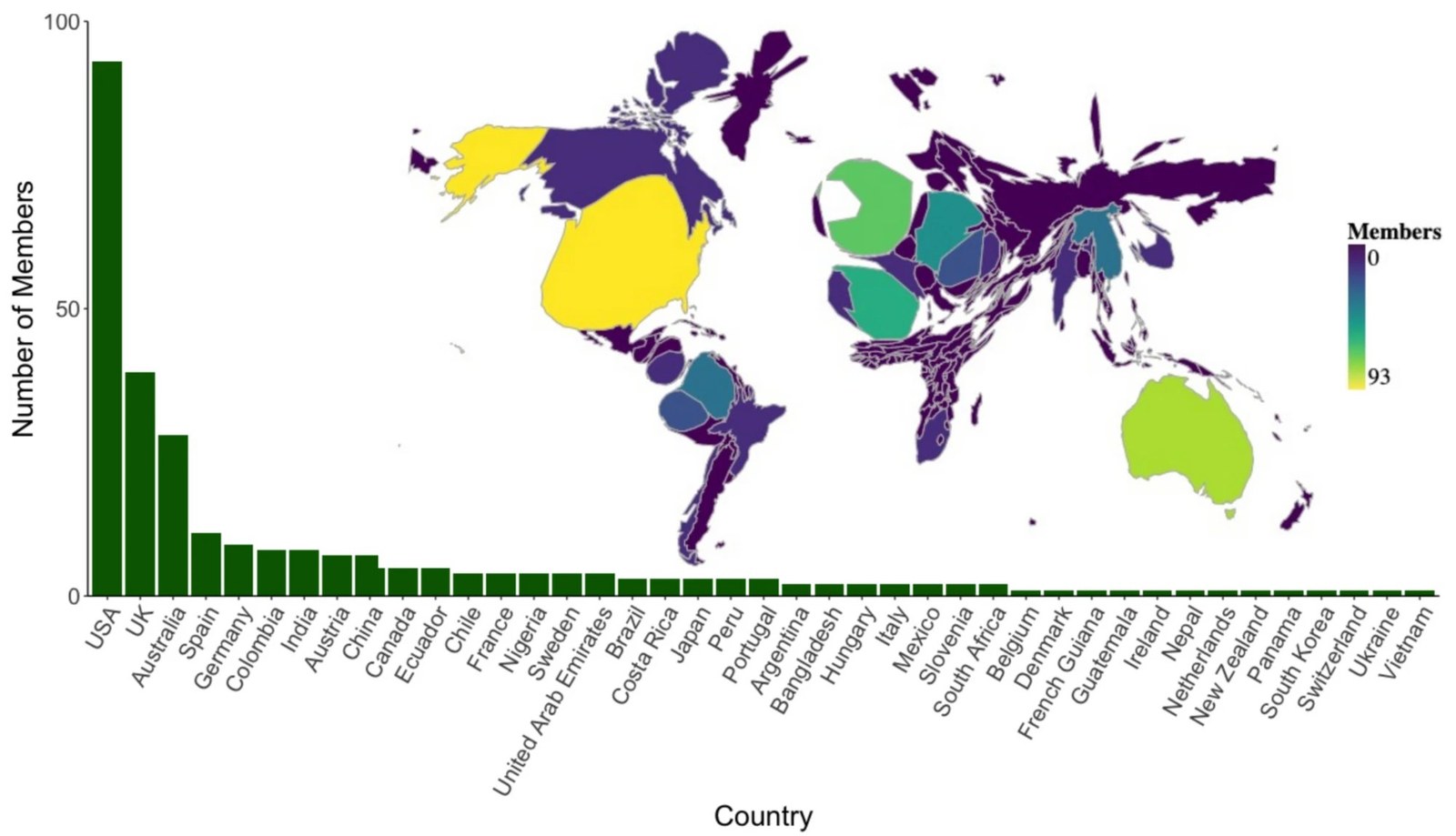
Our Research
We employ computationally intense approaches to fundamental questions in biology — from the patterns of adaptation encoded in molecular sequences to the deep evolutionary history of animals and the evolution of animal systems and traits.
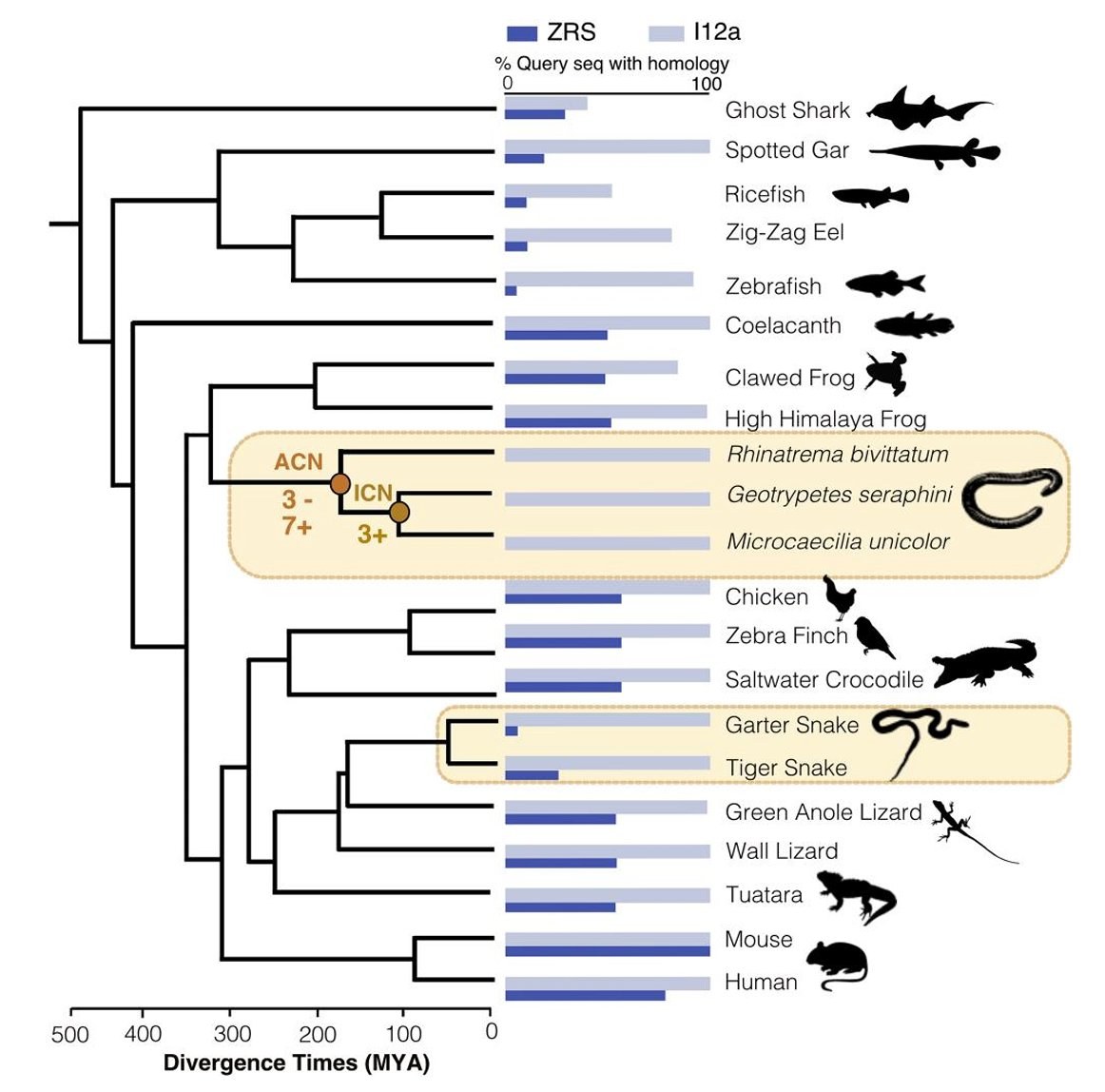 Molecular Evolution
Molecular Evolution
 Phylogenomics
Phylogenomics
 Vertebrate Genomics
Vertebrate Genomics
Phylogenetics & Phylogenomics
Uncovering the tree-like and non-tree-like processes that contribute to phenotypic diversity and speciation. Developing and implementing new methods for constructing genome-scale phylogenetic histories.
Comparative Genomic Zoology & Regulatory Evolution
Investigating the molecular evolution of regulatory networks and the control of gene expression as a primary driver of animal diversity. We study how changes in regulatory architecture — from transcription factor binding sites to non-coding RNA repertoires — underpin the development and evolution of animal organs and systems.
Eukaryotic Origins
Understanding the deep history of the eukaryotic cell, including the hybrid nature of the Eukaryota and the contributions of the Asgard archaea to our understanding of cellular evolution.